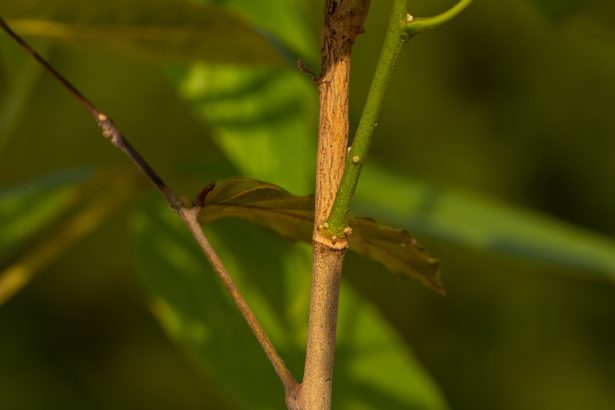
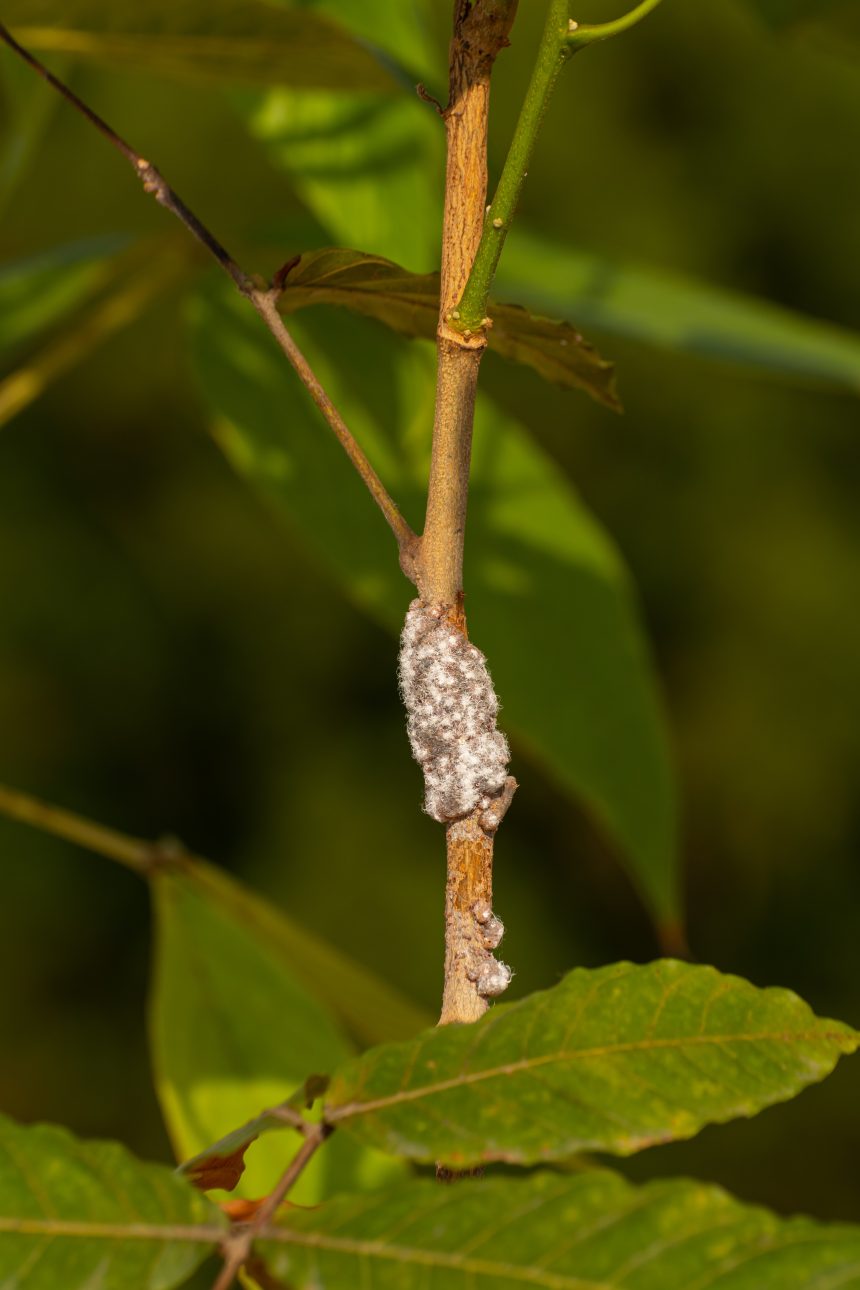
A fungal origin for coveted lac pigment
The colourful pigment extracted from the lac insect may actually be produced by a symbiotic yeast-like organism living inside the…
Fast Four Quiz: Precision Medicine in Cancer
Bacteria ‘nanowires’ could help scientists develop green electronics
Engineered protein filaments originally produced by bacteria have been modified by scientists…
Nuisance Seaweed Found to Produce Compounds with Biomedical Potential
A seaweed considered a threat to the healthy growth of coral reefs…
Slowdown of global warming fleeting
The recent slowdown in the warming rate of the Northern Hemisphere may be a result…
Timing of chemo affects inflammation, mice study suggests
Finding ‘sweet spot’ for drug administration could help patients The time of day that breast…
Organic electronics could lead to cheap, wearable medical sensors
UC Berkeley engineers have created a pulse oximeter sensor composed of all-organic optoelectronics that uses red and green light. The…
Sign Up for Free
Subscribe to our newsletter and don't miss out on our programs, webinars and trainings.
Mimics human tissue, fights bacteria: new biomaterial hits the sweet spot
A new lab-made substance mimics human tissue and could reduce or replace the use of animal-derived materials in biomedical research.…
Your one-stop resource for medical news and education.
Nuisance Seaweed Found to Produce Compounds with Biomedical Potential
A seaweed considered a threat to the healthy growth of coral reefs in Hawaii may possess the ability to produce…
Nuisance Seaweed Found to Produce Compounds with Biomedical Potential
A seaweed considered a threat to the healthy growth of coral reefs in Hawaii may possess the ability to produce…
Cotton-Based Hybrid Biofuel Cell Could Power Implantable Medical Devices
A glucose-powered biofuel cell that uses electrodes made from cotton fiber could someday help power…
New 3D printer uses rays of light to shape objects, transform product design
A new 3D printer uses light to transform gooey liquids into complex solid objects in…
New tools used to identify childhood cancer genes
Using a new computational strategy, researchers at UT Southwestern Medical Center have identified 29 genetic…
A fungal origin for coveted lac pigment
The colourful pigment extracted from the lac insect may actually be produced by a symbiotic…
Combination approach could overcome treatment resistance in deadly breast cancer
QIMR Berghofer-led research in collaboration with Australian oncology company, Kazia Therapeutics, has found that combining…
Tailored brain stimulation treatment results give new hope for people with depression
Medical researchers at QIMR Berghofer have achieved a significant milestone in the treatment of depression,…
Game-Changer in Emergency Medicine: New AI Test Flags Sepsis Hours Before Symptoms Worsen
When the body's reaction to an infection goes awry, it can result in sepsis, a…
Perfumes and lotions disrupt how body protects itself from indoor air pollutants
Fragrances and lotions don't just change the way people smell, they actively alter the indoor…
Medical Milestone: Surgeons Perform First-Ever Human Bladder Transplant
In a groundbreaking medical achievement, surgeons at the University of California, Los Angeles (UCLA) Health…
A Downside of Taurine: It Drives Leukemia Growth
A new scientific study identified taurine, which is made naturally in the body and consumed…
How do therapy dogs help domestic abuse survivors receiving support services?
A new exploration of how therapy dogs can create a safe, nonjudgmental environment for survivors…