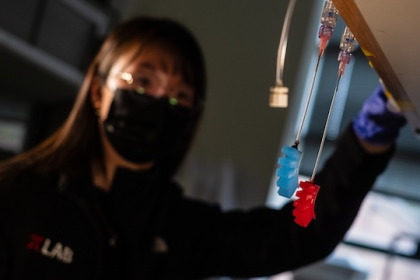
Lab-grown ‘tiny hearts’ bring hope for children and adults with genetic heart disease
Scientists from QIMR Berghofer’s Cardiac Bioengineering Lab have developed lab-grown, three-dimensional heart tissues known as cardiac organoids that mimic the…
Fast Four Quiz: Precision Medicine in Cancer
Nuisance Seaweed Found to Produce Compounds with Biomedical Potential
A seaweed considered a threat to the healthy growth of coral reefs…
Bacteria ‘nanowires’ could help scientists develop green electronics
Engineered protein filaments originally produced by bacteria have been modified by scientists…
Rice research could advance soft robotics manufacturing, design
Quantitative framework enables tunable elastomer curing via temperature control Te Faye Yap, a Rice University…
Elephant seals help uncover slower-than-expected Antarctic melting
Data recorded by elephant seals helped researchers from the…
Glass frogs living near roaring waterfalls wave hello to attract mates
Most frogs emit a characteristic croak to attract the attention of a potential mate. But a few frog species that…
Sign Up for Free
Subscribe to our newsletter and don't miss out on our programs, webinars and trainings.
Retired stars join the young stars’ party in the sky: how evolved stars contribute to the early heating of Earth
Researchers from the University of Sheffield and Imperial College London have spotted a ‘retired’ Asymptotic Giant Branch (AGB) star passing…
Smartphone Apps May Connect to Vulnerable Backend Cloud Servers
Cybersecurity researchers have discovered vulnerabilities in the backend systems that feed content…
Your one-stop resource for medical news and education.
Bacteria ‘nanowires’ could help scientists develop green electronics
Engineered protein filaments originally produced by bacteria have been modified by scientists to conduct electricity. In a study published recently…
Nuisance Seaweed Found to Produce Compounds with Biomedical Potential
A seaweed considered a threat to the healthy growth of coral reefs in Hawaii may possess the ability to produce…
New tools used to identify childhood cancer genes
Using a new computational strategy, researchers at UT Southwestern Medical Center have identified 29 genetic…
New 3D printer uses rays of light to shape objects, transform product design
A new 3D printer uses light to transform gooey liquids into complex solid objects in…
Cotton-Based Hybrid Biofuel Cell Could Power Implantable Medical Devices
A glucose-powered biofuel cell that uses electrodes made from cotton fiber could someday help power…
Lab-grown ‘tiny hearts’ bring hope for children and adults with genetic heart disease
Scientists from QIMR Berghofer’s Cardiac Bioengineering Lab have developed lab-grown, three-dimensional heart tissues known as…
Diver-Operated Microscope Brings Hidden Coral Biology into Focus
The intricate, hidden processes that sustain coral life are being revealed through a new microscope…
A fungal origin for coveted lac pigment
The colourful pigment extracted from the lac insect may actually be produced by a symbiotic…
Combination approach could overcome treatment resistance in deadly breast cancer
QIMR Berghofer-led research in collaboration with Australian oncology company, Kazia Therapeutics, has found that combining…
Tailored brain stimulation treatment results give new hope for people with depression
Medical researchers at QIMR Berghofer have achieved a significant milestone in the treatment of depression,…
Game-Changer in Emergency Medicine: New AI Test Flags Sepsis Hours Before Symptoms Worsen
When the body's reaction to an infection goes awry, it can result in sepsis, a…
Perfumes and lotions disrupt how body protects itself from indoor air pollutants
Fragrances and lotions don't just change the way people smell, they actively alter the indoor…
Medical Milestone: Surgeons Perform First-Ever Human Bladder Transplant
In a groundbreaking medical achievement, surgeons at the University of California, Los Angeles (UCLA) Health…